DNA Synthesis. DNA Polymerases. Enzymes that replicate DNA using a DNA template. DNA polymerases. However, there are also enzymes that. DNA using an RNA template (reverse transcriptases).
DNA without using a template (terminal. Most organisms have more than one type of DNA polymerase. E. Polymerization occurs only 5' to 3'2. Polymerization requires a template to copy: the complementary. Polymerization requires 4 d. NTPs: d. ATP, d. GTP, d.
CTP, d. TTP (TTP. Polymerization requires a pre- existing primer from. The primer is RNA in most organisms, but it. DNA in some organisms; very rarely the primer is a protein. Some examples of DNA polymerases Enzyme Template Primer Other activities Other features E. There are approximately 4. The enzyme is a single large protein with a molecular weight.
Enzyme That Synthesizes Rna From Dna Template In Pcr
Da (1. 03,0. 00 grams per mole). The enzyme. requires a divalent cation (Mg++) for activity and has three enzymatic. DNA Polymerase activity. Proofreading. activity)3. Nick translation activity)The location of the three enzymatic activities. I into two protein fragments (see Klenow fragment of.
I in Table above; both the polymerization and 3'- to- 5' exonuclease. Klenow fragment of pol. I, and the 5'- to- 3'. Like all known DNA polymerases, DNA polymerase I requires a primer.
DNA in the 5' to 3' direction using the complementary strand. The template provides the. Watson- Crick base pairing(Figure. Only the alpha phosphate. NTP is incorporated into newly synthesized DNAThe rate of DNA synthesis by pol I is only.
Transcription (RNA Synthesis). DNA template; The enzyme that performs transcription is referred to. ASSAY FOR RNA SYNTHESIS; Reagents; DNA template, RNA. Transcription is nothing but the formation of RNA from DNA template. Transcription / RNA Synthesis. On protein synthesis. Telomerase: An RNP Enzyme Synthesizes DNA. The demonstration that an RNA moiety provides the template for polymerization of telomeric DNA repeats.
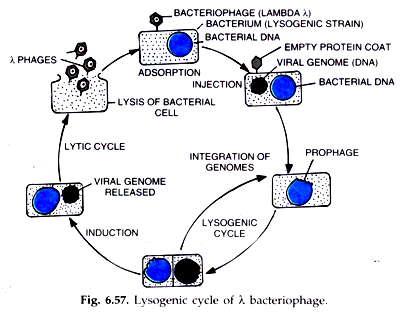
E. In 1. 96. 9, John. Cairns isolated a viable mutant (pol. A) lacking DNA polymerase. I activity, an indication that pol. I is NOT the main enzyme. DNA. Interestingly, pol. A mutants are defective.
As transcription proceeds. The main enzyme responsible for synthesis of DNA from an RNA template is called reverse. RNA polymerase synthesizes an RNA strand. The enzyme that synthesizes messenger RNA from a DNA. DNA from an RNA template. The enzyme that synthesizes messenger RNA from a. Start studying DNA, RNA, and Protein Synthesis. RNA by using one strand of a DNA molecule as a template.
DNA damage. DNA polymerase I also has two additional activities. This was first discovered in .
Enzyme That Synthesizes Rna From Dna Template And Coding
In general, the 3'- to- 5'. DNA synthesis. by a factor of 1.
Thus, when the 5' to 3' polymerization. DNA, the 3'- . to 5' exonuclease proofreading activity immediately removes it. This enzymatic. activity plays an essential role by removing RNA primers during replication. This process. is similar to nick translation (here translation means .
The following points should. Only . This. is called shotgun sequencing. Polymerase Chain Reaction (PCR)The polyerase chain reaction (PCR) is.
DNA. A few uses of PCR are: (a) amplification of desired genomic. DNA sequences with or without subsequent cloning, (b) direct sequencing.
DNA without the need to clone the molecule, (c) medical. DNA fingerprinting), (d) mutating DNA. DNAs. Kary. Mullis, the inventor of the technique was awarded the Nobel prize. Since the introduction of the technique.
The PCR process is based on the inherent. DNA replication. DNA (typically genomic DNA)2. Thermostable DNA polymerase (for example, Taq polymerase)3. Primers (oligonucleotides complementary to specific sequences.
DNA)4. Requires two primers. Amplifies only sequences between two primers.
Surrounding. sequences are not amplified. In vitro DNA replication is 1. DNA replication. Cloning c. DNA (a DNA copy of messenger RNA)Cloned eukaryotic genes cannot be expressed. We will learn about introns. It is possible, however, to make a DNA copy of messenger. RNA, a so called c.
DNA. A cloned c. DNA can be expressed in bacteria. DNA is constructed next to a bacterial promoter. The synthesis. of c. DNA requires the enzyme reverse transcriptase. This. enzyme has the very useful property of making DNA from an RNA.
Eukaryotic messenger RNAs (m. RNAs) are. usually present in cells in low amounts, and must be purified. RNAs (r. RNAs) and. RNAs (t. RNAs). Most eukaryotic m. RNAs are polyadenylated. A nucleotides to the 3'- ends of eukaryotic m.
RNA molecules. The poly. A tails of the m. RNAs will base pair with. T, and . The. r. RNAs and t.
RNAs have no long stretches of A that can base pair. T, and therefore do not stick to the column. After. the r. RNAs and t. RNAs are washed through the column, the purified. RNAs are finally removed by breaking the hydrogen bonding between. A and oligo d. T. Purified m. RNAs can now be converted to c.
DNAs. using reverse transcriptase and oligo d. T as a primer (Figure. BI). The oligo d. T base pairs to. the poly. A tails of the m.
RNAs, and reverse transcriptase synthesizes. DNA (c. DNA) strand using the RNA as a template. The single- stranded DNA can be converted to double- stranded. DNA using random hexamer primers (synthetic DNA strands containing.
DNA polymerase. There are several directions that can be taken. The c. DNA molecules can be cloned. DNA library. A c. DNA encoding a particular.
DNA library with a. Alternatively, the c. DNA mixture. can be amplified by PCR using . The amplified c. DNA product will encode a particular protein. DNA manipulations. This. last method is called RT- PCR (reverse transcriptase.
Transcription and RNA polymerase - An Introduction to Genetic Analysis. As described earlier, transcription relies on the complementary pairing of bases. The. two strands of the double helix separate locally, and one of the separated strands. Next, free nucleotides are aligned on the DNA template by their. The free ribonucleotide A aligns with T in the. DNA, G with C, C with G, and U with A. The process is catalyzed by the enzyme.
RNA polymerase, which attaches and moves. DNA adding ribonucleotides in the growing RNA as shown in Figure 1. Hence, already we see the two. RNA polymerase) in action. Because of the antiparallel nature of the nucleotide pairing, the.
RNA is synthesized 5. RNA polymerase. In most prokaryotes, a single RNA polymerasespecies transcribes all types of. RNA. Figure 1. 0- 7 shows the structure of. RNA polymerase from E. We can see that the enzyme.
The complete enzyme with . The core enzyme contains two . The. addition of the .
The bases are aligned according to homologies, or. Figure. 1. 0- 8). These regions have been termed. At the bottom of Figure. The enzyme then unwinds DNA and begins the. RNA molecule. The dissociative subunit of RNA polymerase, the . First, the holoenzyme.
Figure 1. 0- 9a). The. resulting structure is termed a closed promoter complex (Figure 1. Then, the enzyme binds more. When the bound polymerase causes. DNA duplex, it is said to form an open. Figure. 1. 0- 9c).
This initiation step, the formation of an open complex. Transcription begins, (more..)Elongation. Shortly after initiating transcription, the sigma factor dissociates from the RNA polymerase. The RNA is always synthesized in the 5. The following equation represents the addition of. The energy for the reaction is derived from splitting the high- energy.
PPi), as shown in Figure. Figure 1. 0- 1. 1 gives a. Note how a “transcription bubble” must be. Certain sequences. The sequential addition of nucleotides takes place one at a time in the. The chain grows by the formation of a bond between.
An RNA strand is synthesized in the. Snustad, Principles of. Genetics,(more..)Termination. RNA polymerase also recognizes signals for chain termination, which includes the. RNA and the enzyme from the template.
There are two major. E. The terminator sequences. GC- rich stretch that is followed by a run of.
A’s on the template strand. The corresponding GC sequences on the. RNA are so arranged that the transcript in this region is able to form. Figure 1. 0- 1. 2. The resulting. double- stranded RNA section is called a hairpin loop.
It is. followed by the terminal run of U’s that correspond to the A residues on the DNA. The hairpin loop and section of U residues appear to serve as a signal. RNA polymerase and termination of transcription. A model for rho- dependent termination is shown in. Figure 1. 0- 1. 3. Rho is a hexamer.
ATP to ADP and. Pi drives the termination reaction. The first step in termination.
RNA termed rut. (Figure 1. After. binding, rho pulls the RNA off the RNA polymerase, probably by translocating. RNA, as depicted in Figure. The rut sites are located just.
 RSS Feed
RSS Feed
